ahds¶
Table of Contents
Overview¶
ahds is a Python package to parse and handle Amira (R) files.
It was developed to facilitate reading of Amira (R) files as part of the EMDB-SFF toolkit.
Note
Amira (R) is a trademark of Thermo Fisher Scientific. This package is in no way affiliated with with Thermo Fisher Scientific.
Installation¶
Presently, ahds only works with Python 2.7 but will soon work on Python 3. Please begin by
installing numpy<1.16 using
pip install numpy<1.16
because it is needed to run setup.py. Afterwards you may run
pip install ahds
License¶
Copyright 2017 EMBL - European Bioinformatics Institute
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing,
software distributed under the License is distributed on an
"AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND,
either express or implied. See the License for the specific
language governing permissions and limitations under the License.
Future Plans¶
- Write out valid Amira (R) files
Background and Definitions¶
ahds presently handles two types of Amira (R) files:
- AmiraMesh files, which typically but not necessarily have a
.amextension, and - HyperSurface files, which have
.surfand represent an older filetype.
Both file types consist of two parts:
- a header, and
- one or more data streams.
Headers are structured in a modified VRML-like syntax and differ between AmiraMesh and HyperSurface files in some of the keywords used.
A data stream is a sequence of encoded bytes either referred to in the header by some delimiter (usually @<data_stream_index>, where <data_stream_index> is an integer) or a set of structural keywords (e.g. Vertices, Patches) expected in a predefined sequence.
Headers in Detail¶
AmiraMesh and HyperSurface headers can be divided into four main sections:
- designation
- definitions
- parameters, and
- data pointers.
The designation is the first line and conveys several important details about the format and structure of the file such as:
- filetype (either
AmiraMeshorHyperSurface) - dimensionality (
3D) - format (
BINARY-LITTLE-ENDIAN,BINARYorASCII) - version (a decimal number e.g.
2.1 - extra format data e.g.
<hxsurface>specifying that an AmiraMesh file will contain HyperSurface data
A series of definitions follow that refer to data found in the data pointer sections that either begin with the word ‘define’ or have ‘n’ prepended to a variable. For example:
define Lattice 862 971 200
or
nVertices 85120
This is followed by grouped parameters enclosed in a series of braces beginning with the word ‘Parameters’. Various parameters are then enclosed each beginning with the name of that group of parameters e.g. ‘Materials’
Parameters {
# grouped parameters
Material {
# the names of various materials with attributes
Exterior {
id 0
}
Inside {
id 1,
Color 0 1 1,
Transparency 0.5
}
}
Patches {
# patch attributes
InnerRegion “Insideâ€,
OuterRegion “Exteriorâ€,
BoundaryID 0,
BranchingPoints 0
}
# inline parameters
GridSize <value>,
…
}
The most important set of parameters are materials as these specify colours and identities of distinct segments/datasets within the file.
Finally, AmiraMesh files list a set of data pointers that point to data labels within the file together with additional information to decode the data. We refer to these as data streams because they consist of continuous streams of raw byte data that need to be decoded. Here is an example of data pointers that refer to the location of 3D surface primitives:
Vertices { float[3] Vertices } @1
TriangleData { int[7] Triangles } @2
Patches-0 { int Patches-0 } @3
These refer to three raw data streams each found beginning with the delimiter @<number>. Data stream one (@1) is called Vertices and consists of float triples, two is called TriangleData and has integer 7-tuples and three called Patches- is a single integer (the number of patches). In some cases the data pointer contains the data encoding for the corresponding data pointer.
Lattice { byte Labels } @1(HxByteRLE,234575740)
which is a run-length encoded data stream of the specified length, while
Lattice { byte Data } @1(HxZip,919215)
contains zipped data of the specified length.
Data Streams in Detail¶
AmiraMesh data streams are very simple. They always have a start delimiter made of @ with an index that identifies the data stream. A newline character separates the delimiter with the data stream proper which is either plain ASCII or a binary stream (raw, zipped or encoded).
HyperSurface data streams structured to have the following sections:
# Header
Vertices <nvertices>
# vertices data stream
NBranchingPoints <nbranching_points>
NVerticesOnCurves <nvertices_on_curves>
BoundaryCurves <nboundary_curves>
Patches <npatches>
{
InnerRegion <inner_region_name>
OuterRegion <outer_region_name>
BoundaryID <boundary_id>
BranchingPoints <nbranching_points>
Triangles <ntriangles>
# triangles data stream
} # repeats for as <npatches> times
HyperSurface data streams can be either plain ASCII or binary.
ahds Modules¶
ahds has three main modules:
ahds.grammarspecifies an EBNF grammarahds.headerahds.data_stream
These modules are tied into a user-level class called ahds.AmiraFile that does all the work for you.
>>> from ahds import AmiraFile
>>> # read an AmiraMesh file
>>> af = AmiraFile('am/test7.am')
>>> af.header
<AmiraHeader with 4 bytes>
>>> # empty data streams
>>> af.data_streams
>>> print af.data_streams
None
>>> # we have to explicitly read to get the data streams
>>> af.read()
>>> af.data_streams
<class 'ahds.data_stream.DataStreams'> object with 13 stream(s): 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13
>>> for ds in af.data_streams:
... print ds
...
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 2,608 bytes
# we get the n-th data stream using the index/key notation
>>> af.data_streams[1].encoded_data
'1 \n2 \n3 \n'
>>> af.data_streams[1].decoded_data
[1, 2, 3]
>>> af.data_streams[2].encoded_data
'69 \n120 \n116 \n101 \n114 \n105 \n111 \n114 \n0 \n73 \n110 \n115 \n105 \n100 \n101 \n0 \n109 \n111 \n108 \n101 \n99 \n117 \n108 \n101 \n0 \n'
>>> af.data_streams[2].decoded_data
[69, 120, 116, 101, 114, 105, 111, 114, 0, 73, 110, 115, 105, 100, 101, 0, 109, 111, 108, 101, 99, 117, 108, 101, 0]
>>> # read an HyperSurface file
>>> af = AmiraFile('surf/test4.surf')
>>> af.read()
>>> af.data_streams
<class 'ahds.data_stream.DataStreams'> object with 5 stream(s): Patches, NBranchingPoints, BoundaryCurves, Vertices, NVerticesOnCurves
# HyperSurface files have pre-set data streams
>>> af.data_streams['Vertices'].decoded_data[:10]
[(560.0, 243.0, 60.96875), (560.0, 242.9166717529297, 61.0), (559.5, 243.0, 61.0), (561.0, 243.0, 60.95833206176758), (561.0, 242.5, 61.0), (561.0384521484375, 243.0, 61.0), (559.0, 244.0, 60.94444274902344), (559.0, 243.5, 61.0), (558.9722290039062, 244.0, 61.0), (560.0, 244.0, 60.459999084472656)]
ahds.grammar¶
This module describes the header grammar for Amira (R) (AmiraMesh and HyperSurface) files and so depends on simpleparse Python package. It defines a single class (ahds.grammar.AmiraDispatchProcessor) and four functions.
ahds.grammar.AmiraDispatchProcessor is a subclass of simpleparse.dispatchprocessor which implements the core functionality required to use the grammar. Each grammar token has a corresponding method defined on this class which determines how the data associated with that token will be rendered. Data can be rendered as a single or multimap, string, number, or in custom format.
ahds.grammar.get_parsed_data(fn, *args, **kwargs)()is the user-level function that takes a filename and returns structured parsed data. It depends on the other three functions defined:ahds.grammar.detect_format(fn, format_bytes=50, verbose=False)()returns eitherAmiraMeshorHyperSurfacegiven a file name and arguments,ahds.grammar.get_header(fn, file_format, header_bytes=20000, verbose=False)()returns the header portion based on the file format determined by detect_format(…), andahds.grammar.parse_header(data, verbose=False)()converts the raw header data returned byahds.grammar.get_header(...)()into a structured header based on AmiraDispatchProcessor.
ahds.header¶
This module converts the structured header from the ahds.grammar module into an object with the sections of the header (designation, definitions, parameters ``and ``data pointers) and corresponding structured data available as attributes. That is, it converts the header:
# AmiraMesh BINARY-LITTLE-ENDIAN 2.1
define Lattice 862 971 200
Parameters {
Materials {
Exterior {
Id 1
}
Inside {
Color 0.64 0 0.8,
Id 2
}
Mitochondria {
Id 3,
Color 0 1 0
}
Mitochondria_ {
Id 4,
Color 1 1 0
}
mitochondria__ {
Id 5,
Color 0 0.125 1
}
NE {
Id 6,
Color 1 0 0
}
}
Content "862x971x200 byte, uniform coordinates",
BoundingBox 0 13410.7 0 15108.4 1121.45 4221.01,
CoordType "uniform"
}
Lattice { byte Labels } @1(HxByteRLE,4014522)
into an ahds.header.AmiraHeader object.
>>> from ahds.header import AmiraHeader
>>> amira_header = AmiraHeader.from_file('am/test2.am')
>>> amira_header.designation.attrs
['filetype', 'dimension', 'format', 'version', 'extra_format']
>>> amira_header.designation.filetype
'AmiraMesh'
>>> amira_header.designation.dimension
>>> amira_header.designation.format
'BINARY-LITTLE-ENDIAN'
>>> amira_header.definitions.attrs
['Lattice']
>>> amira_header.definitions.Lattice
[862, 971, 200]
>>> amira_header.parameters.attrs
['Materials', 'Content', 'BoundingBox', 'CoordType']
>>> amira_header.parameters.Materials.attrs
['Exterior', 'Inside', 'Mitochondria', 'Mitochondria_', 'mitochondria__', 'NE']
>>> amira_header.parameters.Materials.Exterior.attrs
['Id']
>>> amira_header.parameters.Materials.Exterior.Id
1
>>> amira_header.parameters.Content
'"862x971x200 byte, uniform coordinates",'
>>> amira_header.parameters.BoundingBox
[0, 13410.7, 0, 15108.4, 1121.45, 4221.01]
>>> amira_header.parameters.CoordType
'"uniform"'
>>> amira_header.data_pointers.attrs
['data_pointer_1']
>>> amira_header.data_pointers.data_pointer_1.attrs
['pointer_name', 'data_format', 'data_dimension', 'data_type', 'data_name', 'data_index', 'data_length']
>>> amira_header.data_pointers.data_pointer_1.pointer_name
'Lattice'
>>> amira_header.data_pointers.data_pointer_1.data_format
'HxByteRLE'
>>> amira_header.data_pointers.data_pointer_1.data_dimension
>>> amira_header.data_pointers.data_pointer_1.data_type
'byte'
>>> amira_header.data_pointers.data_pointer_1.data_name
'Labels'
>>> amira_header.data_pointers.data_pointer_1.data_index
1
>>> amira_header.data_pointers.data_pointer_1.data_length
4014522
This module consists of two main classes: ahds.header.AmiraHeader is the user-level class and ahds.header.Block which is a container class for a block of structured data from an Amira (R) header.
AmiraHeader has one constructor: ahds.header.AmiraHeader.from_file(fn, *args, **kwargs)() which takes an Amira (R) file by name and arguments and returns an ahds.header.AmiraHeader object with all attributes set as described above. Alternatively, one can use the initiator form to pass structured data directly: ahds.header.AmiraHeader(parsed_data) which returns an ahds.header.AmiraHeader object configured appropriately.
- The raw data structured data is available as read-only property:
ahds.header.AmiraHeader.raw_header - Internally the
ahds.header.AmiraHeaderclass implements a set of private methods which individually load the four data sections (designation,definitions,parameters, anddata pointers).
The ahds.header.Block class is a container class which converts structured groups to attributes and has two main attributes:
ahds.header.Block.nameprovides the name of the current block
>>> amira_header.designation.name
'designation'
>>> amira_header.parameters.Materials.name
'Materials'
>>> amira_header.parameters.Materials.Exterior.name
'Exterior'
ahds.header.Block.attrsprovides the attributes available on thisahds.header.Block
>>> amira_header.designation.attrs
['filetype', 'dimension', 'format', 'version', 'extra_format']
>>> amira_header.designation.format
'BINARY-LITTLE-ENDIAN'
A given Materials block has two special features:
Block.ids returns the list of ids for all materials. This is important when decoding HxByteRLE compressed data
Block[id] returns the material for the given id using index notation.
>>> amira_header.parameters.Materials.ids
[1, 2, 3, 4, 5, 6]
>>> amira_header.parameters.attrs
['Materials', 'Content', 'BoundingBox', 'CoordType']
# ids attribute is only available for ‘Material’ blocks within ‘parameters’ section
>>> amira_header.parameters.Content.ids
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
AttributeError: 'str' object has no attribute 'ids'
# we can get the name of a material of the given id
>>> amira_header.parameters.Materials[4].name
'Mitochondria_'
ahds.data_stream¶
This is most complex module implementing a hierarchy of classes describing various data streams within Amira (R) files. It has 22 classes and five functions
Classes¶
There are three categories of classes:
- A user-level class that encapsulates (2) below.
- Classes describing Amira (R) data streams
- Classes describing AmiraMesh data streams
- Classes describing HyperSurface data streams
- Data conversion classes (AmiraMesh only)
- Classes abstracting images
- Classes abstracting contours
The user-level ahds.data_stream.DataStreams class is the preferred way to use the module. It takes the name of an Amira (R) file and encapsulates an iterator of data streams.
>>> from ahds import data_stream
>>> data_streams = data_stream.DataStreams('am/test6.am')
>>> data_streams
<class 'ahds.data_stream.DataStreams'> object with 2 stream(s): 1, 2
>>> for ds in data_streams:
... print ds
...
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 968,909 bytes
<class 'ahds.data_stream.AmiraMeshDataStream'> object of 968,909 bytes
Functions¶
The functions implemented in this module decode data streams.
ahds.data_stream.hxbyterle_decode()decodesHxByteRLEdata streamsahds.data_stream.hxzip_decode(data_size, data)()unzips zlib-compressed data streamsahds.data_stream.unpack_binary(data_pointer, definitions, data)()unpacks the structured data stream according to the attributes specified in the data’s data pointerahds.data_stream.unpack_ascii(data)()converts rows of ASCII data into numerical data
Classes in Detail¶
DataStreams class¶
The following attributes are available on objects of this class:
ahds.data_stream.DataStreams.file- filename of Amira (R) fileahds.data_stream.DataStreams.header- an object of classahds.header.AmiraHeaderencapsulating the header data in four sections (designation,definitions,parameters, anddata pointers)ahds.data_stream.DataStreams.filetype- the filetype as specified in (ii) above.ahds.data_stream.DataStreams.stream_data- all raw data from the file (including the header)len(DataStreams)- the number of data streams containedahds.data_stream.DataStreams[<index>]- returns the data stream of the index specified (as defined in the data_pointers section of the header object
Classes describing Amira (R) data streams¶
The following diagrams illustrates the hierarchy of classes:
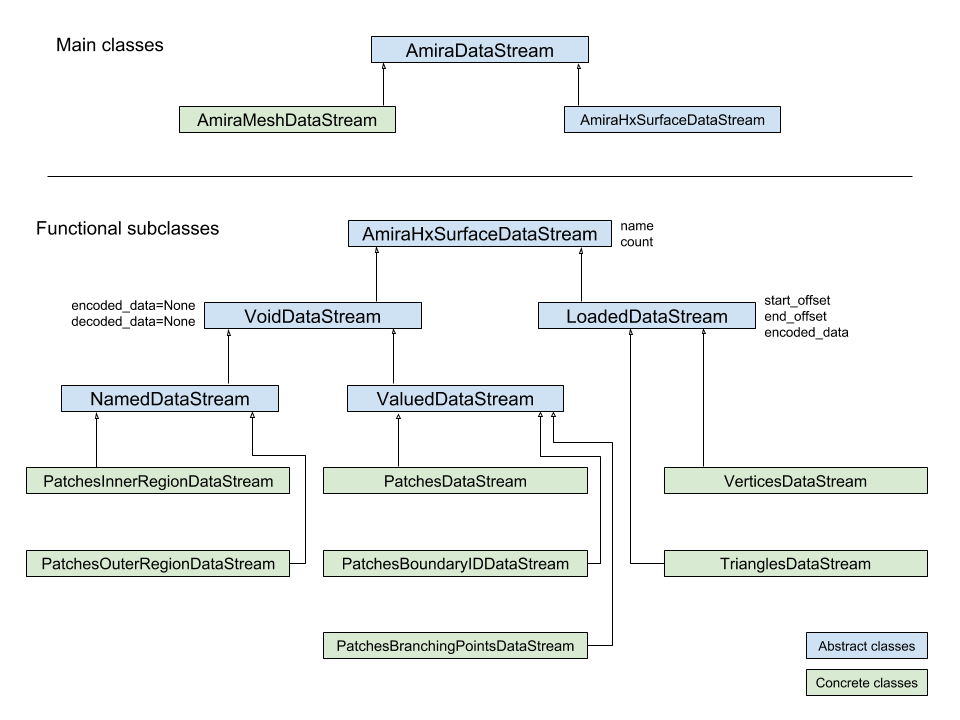
Classes describing Amira (R) data streams
ahds.data_stream.AmiraDataStreamis the base class for all data stream classes and defines the following attributes:
ahds.data_stream.AmiraDataStream.header- anahds.header.AmiraHeaderobjectahds.data_stream.AmiraDataStream.data_pointer- theahds.header.AmiraHeader.data_pointers.data_pointer_Xfor this data streamahds.data_stream.AmiraDataStream.stream_data- the raw file dataahds.data_stream.AmiraDataStream.encoded_data- the encoded data for this stream;NoneforVoidDataStreamsubclassesahds.data_stream.AmiraDataStream.decoded_data- the decoded data for this stream;NoneforVoidDataStreamsubclassesahds.data_stream.AmiraDataStream.decoded_length- the number of items (tuples, integers) in decoded data
The two main subclasses of ahds.data_stream.AmiraDataStream are ahds.data_stream.AmiraMeshDataStream, which is a concrete class representing all AmiraMesh data streams, and ahds.data_stream.AmiraHxSurfaceDataStream, which abstractly defines HyperSurface data streams.
There are two main AmiraHxSurfaceDataStream subclasses:
ahds.data_stream.VoidDataStreamrepresentsahds.data_stream.AmiraHxSurfaceDataStreamdata streams that only have a name and value but no actual encoded data (on the following line). There are two subclasses:
ahds.data_stream.NamedDataStreamsubclasses have a strings after data stream name. The two concrete subclasses are:
ahds.data_stream.PatchesInnerRegionDataStreamfor the name of an inner region of a patch (seePatchesDataStream), andahds.data_stream.PatchesOuterRegionDataStreamfor corresponding name of the outer region of a patch.
ahds.data_stream.ValuedDataStreamhave an integer value after the data stream name. The three concrete subclasses are:
ahds.data_stream.PatchesBoundaryIDDataStreamhold the boundary ID of a patch,ahds.data_stream.PatchesBranchingPointsDataStreamstores the number of branching points, andahds.data_stream.PatchesDataStreamwith the number of patches, which is a specialahds.data_stream.ValueDataStreamthat contains an iterable of patches each containing aPatches<X>DataStreamobjects.
ahds.data_stream.LoadedDataStreamrepresentahds.data_stream.AmiraHxSurfaceDataStreamdata streams that have a name, a value and encoded data. The two main concrete subclasses are:
ahds.data_stream.VerticesDataStreamrepresents data streams with float-triples, andahds.data_stream.PatchesTrianglesDataStreamrepresents data streams within a patch with triples of 1-based indices (triangles) of vertices specified in theahds.data_stream.VerticesDataStream.
Conversion classes¶
There are two groups of conversion classes which only apply to (some) AmiraMesh data streams: Conversion classes
- Image conversion classes consist of a image container class
ahds.data_stream.ImageSetand anahds.data_stream.Imageclass. ImageSet objects that can be iterated to giveahds.data_stream.Imageobjects are returned from theahds.data_stream.AmiraMeshDataStream.to_images()method call.
>>> # decode the data stream to images
>>> images = ds[1].to_images()
>>> images
<ImageSet with 200 images>
>>> for image in images:
... print image
...
<Image with dimensions (971, 862)>
<Image with dimensions (971, 862)>
<Image with dimensions (971, 862)>
...
<Image with dimensions (971, 862)>
<Image with dimensions (971, 862)>
- Contour conversion classes convert individual images into sets of contours (
ahds.data_stream.ContourSet) iterable as individualahds.data_stream.Contoursobjects. They are obtained from calls to theahds.data_stream.Image.as_contoursproperty. Furthermore, theahds.data_stream.Image.as_segmentsproperty call returns a dictionary of the correspondingahds.data_stream.ContourSetobject indexed by the z plane.
>>> # contours per image
>>> # the dictionary key is the Amira Id for the segment (the Id of the Material)
>>> # a segment can have several non-overlapping contours (or polylines)
>>> for image in images:
... print image.as_contours
...
{2: <class 'ahds.data_stream.ContourSet'> with 15 contours, 3: <class 'ahds.data_stream.ContourSet'> with 3 contours, 5: <class 'ahds.data_stream.ContourSet'> with 2 contours}
{2: <class 'ahds.data_stream.ContourSet'> with 18 contours, 3: <class 'ahds.data_stream.ContourSet'> with 3 contours, 5: <class 'ahds.data_stream.ContourSet'> with 2 contours}
...
{2: <class 'ahds.data_stream.ContourSet'> with 15 contours, 3: <class 'ahds.data_stream.ContourSet'> with 1 contours, 5: <class 'ahds.data_stream.ContourSet'> with 3 contours}
{2: <class 'ahds.data_stream.ContourSet'> with 15 contours, 3: <class 'ahds.data_stream.ContourSet'> with 1 contours, 5: <class 'ahds.data_stream.ContourSet'> with 3 contours}
>>> # separate individual segments
>>> images.segments
{1: {110: <class 'ahds.data_stream.ContourSet'> with 1 contours}, 2: {0: <class 'ahds.data_stream.ContourSet'> with 15 contours, 1: <class 'ahds.data_stream.ContourSet'> with 18 contours, ..., 198: <class 'ahds.data_stream.ContourSet'> with 3 contours, 199: <class 'ahds.data_stream.ContourSet'> with 3 contours}}